Selected publications
- Inter-tissue convergence of gene expression during ageing suggests age-related loss of tissue and cellular identity
- Temporal changes in the gene expression heterogeneity during brain development and aging
- The widespread increase in inter-individual variability of gene expression in the human brain with age
Theme 01
Stochasticity & Regulation in Aging
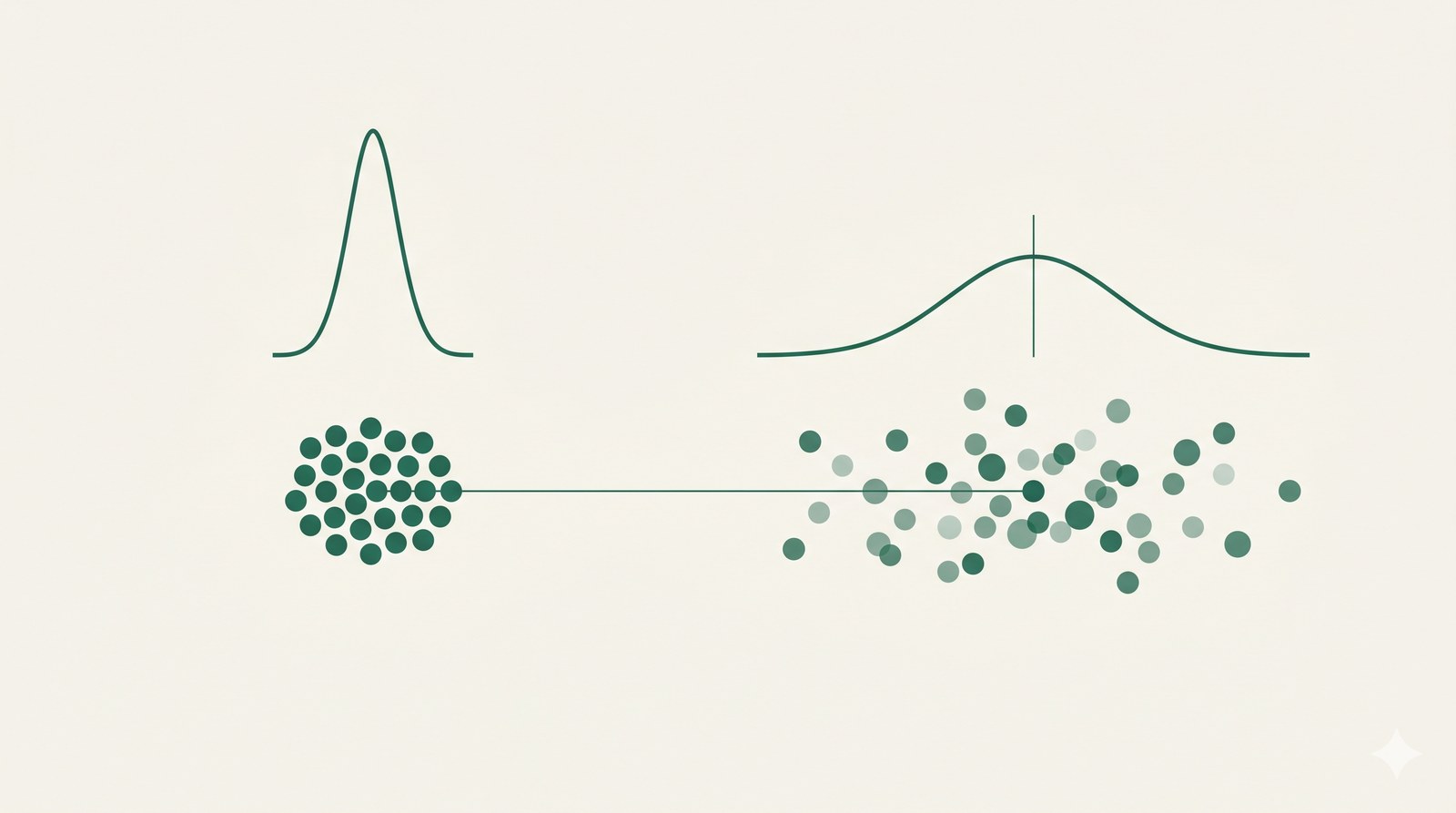
Most aging biomarkers describe what changes on average — average expression, average methylation. But the variance changes too. Regulatory networks lose their tightness with age. Cells of nominally the same type drift apart. Inter-individual differences widen, so that animals of identical chronological age can become noticeably more divergent at the molecular level than they were when young. The noise in the context of ageing is not an artefact; it is a signal.
We are interested in the causes, biological relevance, and intervention potential of this regulatory decay. Is it an upstream driver of age-related dysfunction, or a downstream readout of more proximal failures? Is there a shared geometry of regulatory decay across organisms, tissues, and omics layers — or does each system fail along its own axis? Does the variance signal carry predictive information that mean-based biomarkers miss — for disease risk, drug response, individual trajectories? And is it reversible: do interventions that extend healthspan also restore regulatory coordination, or do they only buy time?